Pili becoming a hot topic in gram-positives
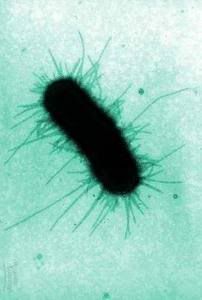 Pili (singular: pilus) are bacterial organelles--thin tubes of protein that function in attachment and bacterial sex, as well as immune evasion. Traditionally, studies of pili have been carried out in gram-negative bacteria, such as E. coli and Neisseria species; very little was known about pili in gram-positives. A few recent high-profile papers have changed that.
Pili (singular: pilus) are bacterial organelles--thin tubes of protein that function in attachment and bacterial sex, as well as immune evasion. Traditionally, studies of pili have been carried out in gram-negative bacteria, such as E. coli and Neisseria species; very little was known about pili in gram-positives. A few recent high-profile papers have changed that.Back in July, an Italian group analyzed the genomic sequences of 2 isolates of Streptococcus agalactiae, or group B streptococcus, in order to identify surface proteins which may be vaccine candidates. (More on other outcomes of similar research here). 3 of the proteins they identified were found to be pilus-like structures extending from the bacterial surface when visualized using immunogold electron microscopy (essentially, using gold particles stuck to antibodies)--the first ever description of pili in these bacteria, and a new target for investigation of virulence factors and vaccine candidates, as immunization with these proteins protected mice from challenge with a lethal dose of the bacteria.
Now, in this week's Proceedings of the National Academy of Sciences (PNAS for the acronymophiles), a similar structure is described in Streptococcus pyogenes, the group A streptococcus. Even more interesting is the finding that several of these pilin proteins seem to be equivalent to the T antigens in S. pyogenes.
A bit of background is in order here. Way back in the days of yore, a typing scheme for S. pyogenes was worked out based on components on the surface of the bacteria. The "A" in group A streptococcus reflects a particular type of carbohydrate this species carries, which is distinct from the "B" carbohydrate of Streptococcus agalactiae, from the "C" carbohydrate of Streptococcus equi and others in that group, etc., etc. So, this is used to differentiate between different species. Once you start investigating differences within a species, there are different methods to type them. In group B strep, they're serotyped using differences in the bacterium's capsule antigens. In group A, they're serotyped using differences in antigenic properties of a surface protein, called the M protein. (Now this is mostly done a step backward, by sequencing the emm gene, which encodes the M protein, rather than dealing with around 100 different antisera to the various M proteins. Cheaper, less messy, and more repeatable between labs). In addition to M typing, the T surface antigen has been used to further distinguish between strains, although much less attention has been paid to this antigen. It looks like that will likely change in short order with this finding.
Anyhoo, back to the paper. I mentioned the Italian group above, who carried out the research on GBS pili. This same group carried out the study on GABHS (group A beta-hemolytic streptococci). They first searched simply on genetic similarity to the GBS (group B streptococci) pili genes, and came up empty. So they took another tactic, looking for groups of closely linked genes coding for surface proteins containing LPXTG motifs (typical of cell surface-associated proteins in gram-positive bacteria) in close proximity to genes coding for variant sortase enzymes.
They went on to characterize the genes they found, expressing them in E. coli and raising antibodies against them. They found similarites between the new proteins in GABHS and the pili they'd recently identified in GBS, and when they analyzed the bacteria by electron microscopy using this antisera, they could see the structures of the pili (panel H is the negative control):

They also used antiserum to various T antigens and determined that there was indeed cross-reactivity between the T antigens and the pilin proteins they'd identified; but it remains unclear whether all T antigens are components of pili, or just the ones they've identified.
Finally, they carried out a mouse immunization experiment, inoculating mice with 3 of the pili proteins, and then challenged with a strain known to be lethal to mice. ~70% survived, which was similar to the survival rate in mice challenged with the M1 protein (vaccines consisting of M protein components have long been suggested and tested, though they remain problematic due to the fact that some M proteins cause autoimmunity).
Definitely lots of interesting work going on in streptococcal genetics and pathogenesis. Now if only someone could make them more amenable to genetic manipulation, we'd be golden.


